
DeltaFold is an interdisciplinary research project at the interface of computational algebraic topology, bioinformatics, and biology, which I co-lead with Céline Brochier-Armanet. The project is supported by the CNRS as a PRIME project and by the ANR through the project ThermAdapt.
The project brings together researchers in mathematics, bioinformatics, biology, and biochemistry to develop new geometric and spectral methods (“Delta”) for analyzing protein structures (“Fold”) and modeling the evolution of life-history traits.
At its core, DeltaFold introduces a mathematical framework to extract evolutionary signals from the three-dimensional structures of proteins. Using topological data analysis, the project builds geometric representations of protein structures based on multilayer simplicial complexes, enabling predictive models that combine persistent homology, spectral geometry, and deep learning.
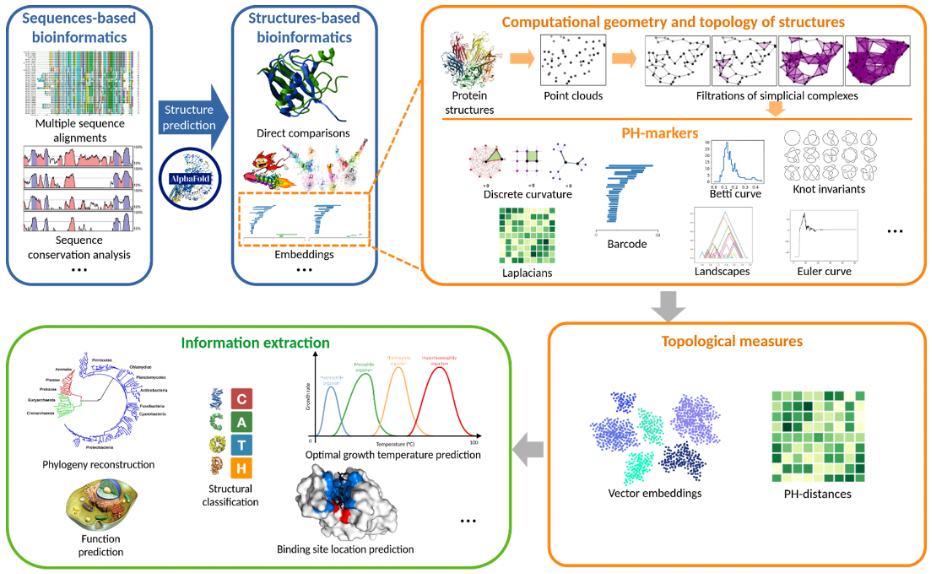
DeltaFold introduces a new approach to decoding biological information from the shape of proteins using topological data analysis (TDA). By translating protein structures into mathematical objects such as simplicial complexes and persistent homology barcodes, TDA captures their global and multiscale geometric features.
These methods track topological features across scales, revealing patterns linked to evolutionary adaptation, functional constraints, and protein stability. Combined with spectral geometry and machine learning, they form a powerful framework to analyze and predict protein structure and function.
Using persistent homology, spectral geometry, and deep learning, DeltaFold develops models that can:
All methods and predictive tools are released as open source through the DeltaFold modeling Python library.